# A tibble: 6 × 9
species date geo_group region losses dead discarded escaped other
<chr> <date> <chr> <chr> <dbl> <dbl> <dbl> <dbl> <dbl>
1 salmon 2020-01-01 area 1 31425 28126 3299 0 0
2 salmon 2020-01-01 area 2 324116 277888 46113 0 115
3 salmon 2020-01-01 area 3 844829 776983 63770 0 4076
4 salmon 2020-01-01 area 4 676852 623159 51823 0 1870
5 salmon 2020-01-01 area 5 109269 97627 11424 0 218
6 salmon 2020-01-01 area 6 548921 531193 15710 0 2018Working smarter with dplyr 1.2.0
R-Ladies Rome | Isabella Velásquez
Introduction

Introduction
⬢ Slides available at: https://ivelasq-dplyr-1-2-0.share.connect.posit.cloud
⬢ Links available at the end of the slide deck


Today’s data
Salmonid Mortality Data from TidyTuesday
⬢ Salmonid mortality datasets published by the Norwegian Veterinary Institute
⬢ Two datasets are shared, the monthly mortality data, and the monthly loses data
⬢ Data from 2020

Today’s data
monthly_losses_data
dplyr
dplyr is a grammar of data manipulation, providing a consistent set of verbs that help you solve the most common data manipulation challenges
Quick review of dplyr functions
arrange() changes the ordering of the rows
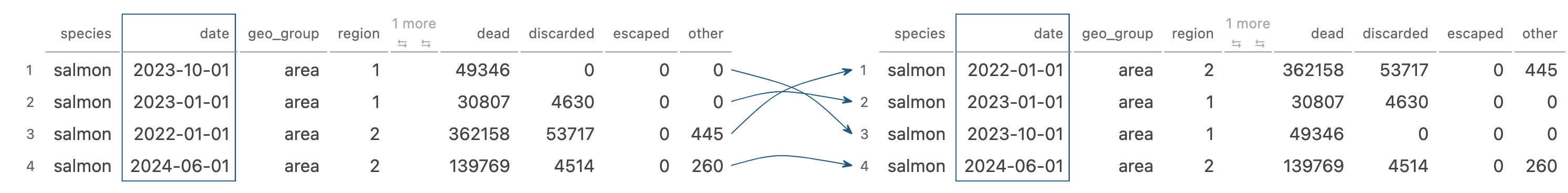
Image generated by Tidy Data Tutor
Quick review of dplyr functions
select() picks variables based on their names
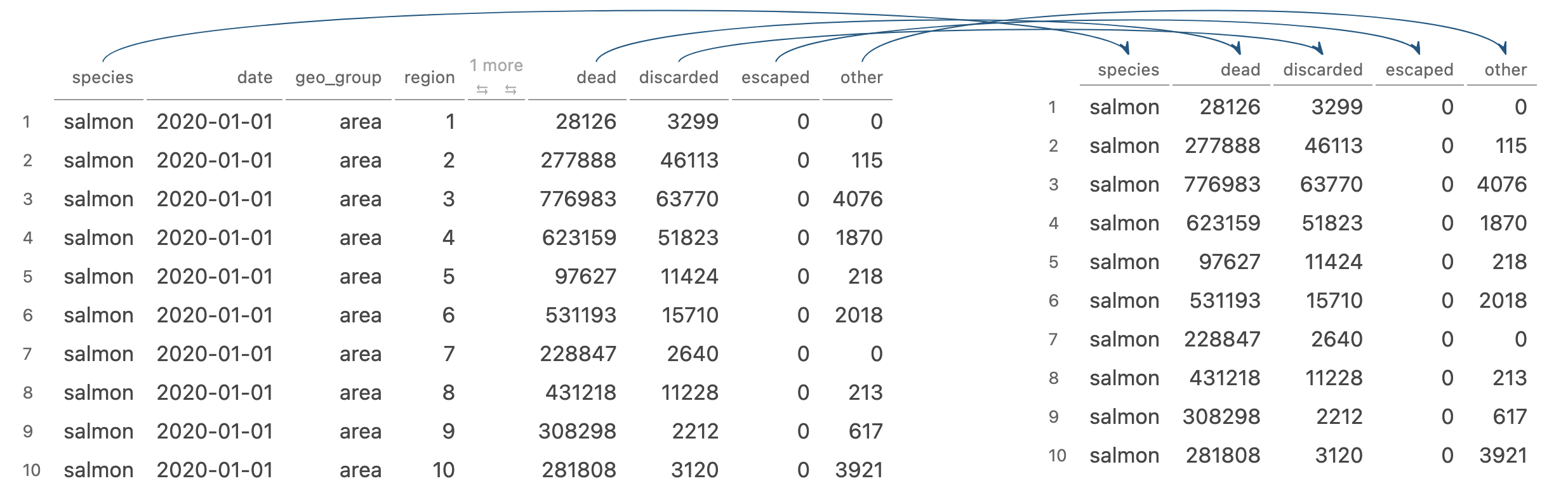
Image generated by Tidy Data Tutor
Quick review of dplyr functions
summarise()/summarize() reduces multiple values down to a single summary
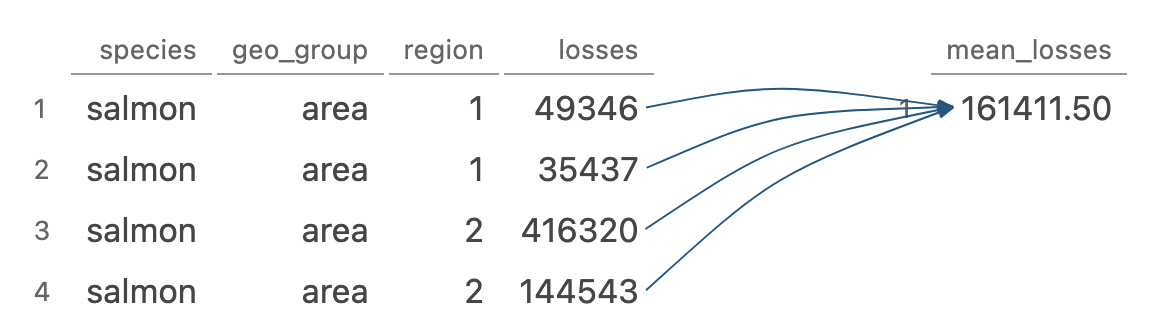

Image generated by Tidy Data Tutor
Quick review of dplyr functions
group_by() allows you to perform any operation “by group”
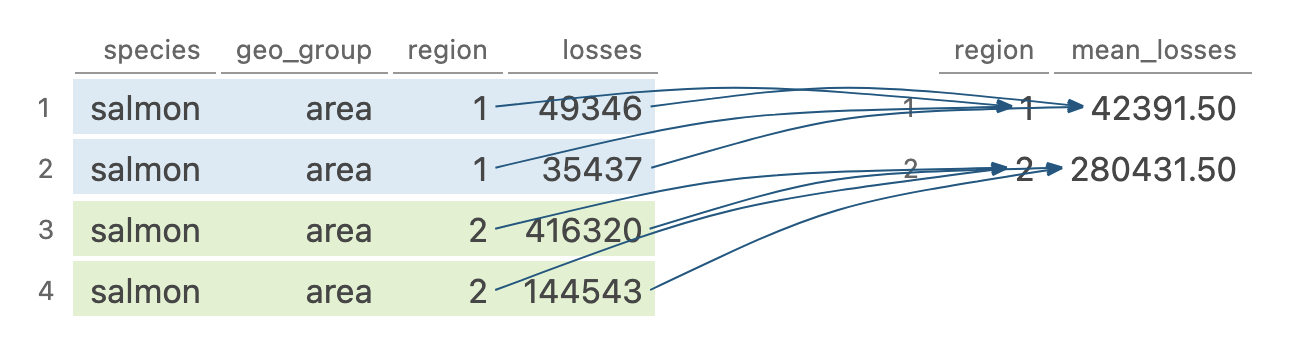

Image generated by Tidy Data Tutor
Quick review of dplyr functions
mutate() adds new variables that are functions of existing variables
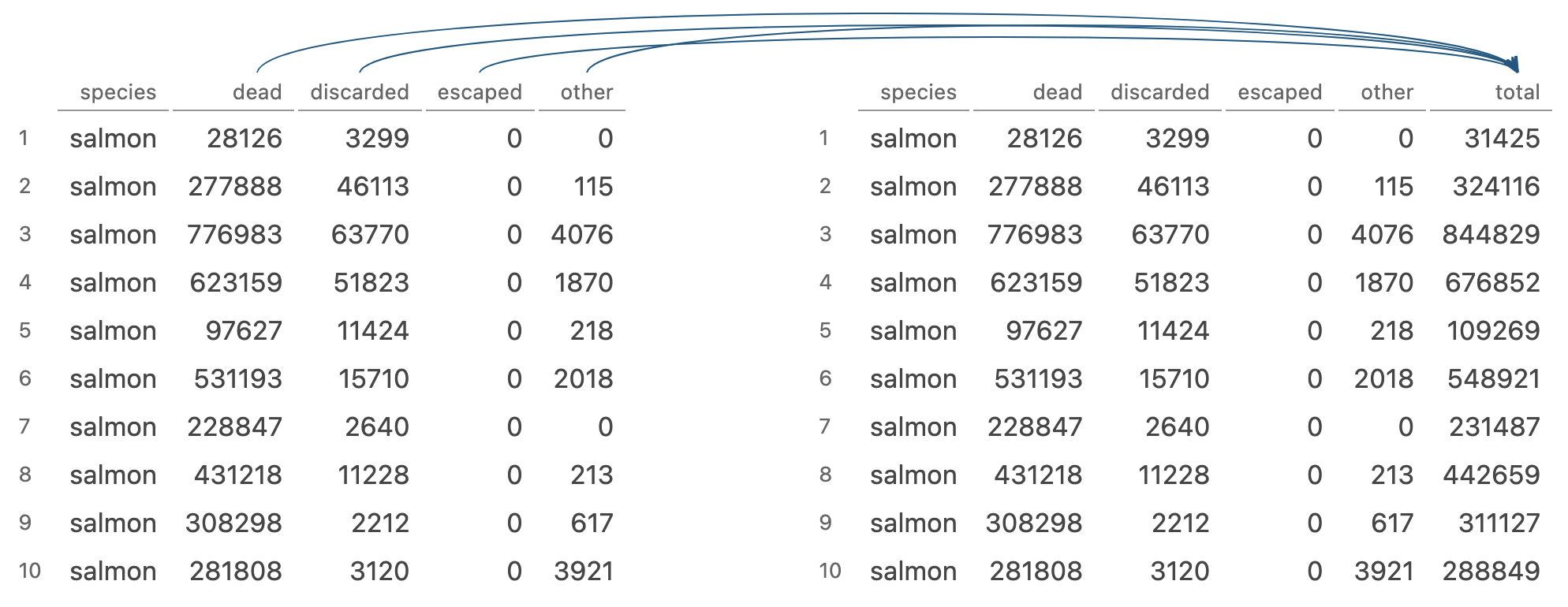
Image generated by Tidy Data Tutor
Quick review of dplyr functions
case_when() checks each condition in order and uses the first match to determine the value of a new variable

Quick review of dplyr functions
filter() picks cases based on their values
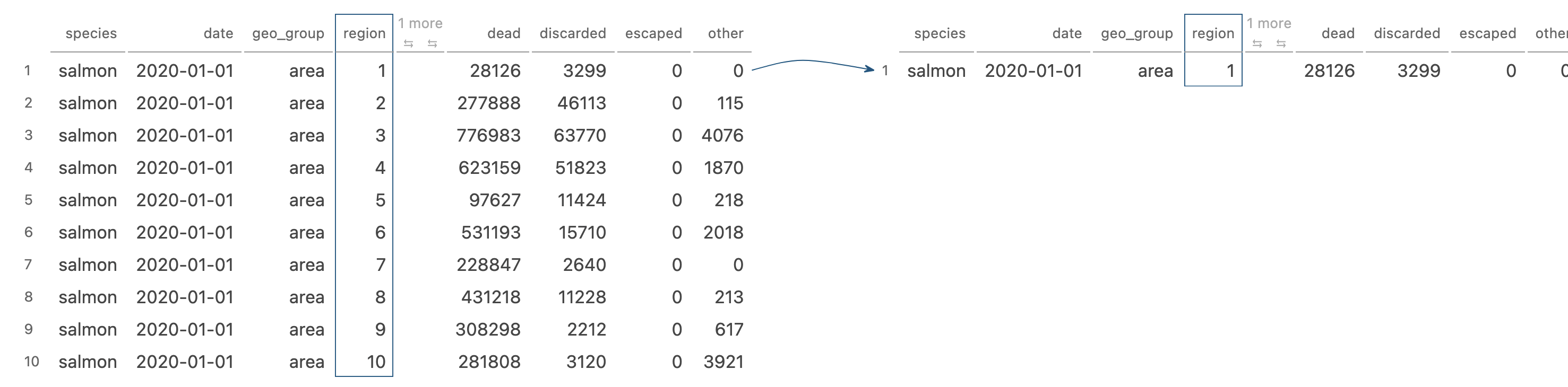
Image generated by Tidy Data Tutor
Quick review of dplyr functions
So many helpful functions!
⬢ distinct()
⬢ slice()
⬢ count()
⬢ pull()
⬢ relocate()
⬢ rename()
⬢ *_join()
⬢ …
But for now, let’s focus on:
⬢ filter()
⬢ mutate() + case_when()
dplyr 1.2.0
filter_out()
The problem with using filter() to exclude
is a little ambiguous! Are you keeping (filtering in) Region 1 or dropping (filtering out) Region 1?
⬢ filter() is optimized for the case of keeping rows, but using it for dropping rows can require complex logic
The problem with using filter() to exclude
Let’s look at this sample dataset:
The problem with using filter() to exclude
Drop rows where region is 3 and losses are greater than 700,000.
# A tibble: 5 × 3
species region losses
<chr> <chr> <dbl>
1 salmon 1 31425
2 salmon 2 324116
3 salmon 3 844829
4 salmon 3 676852
5 salmon 3 NA
The problem with using filter() to exclude
Drop rows where region is 3 and losses are greater than 700,000. (In this case, Row 3)
The problem with using filter() to exclude
Oh no! What happened to our row with NA under losses?
Pre-filter:
# A tibble: 5 × 3
species region losses
<chr> <chr> <dbl>
1 salmon 1 31425
2 salmon 2 324116
3 salmon 3 844829
4 salmon 3 676852
5 salmon 3 NAPost-filter:
# A tibble: 3 × 3
species region losses
<chr> <chr> <dbl>
1 salmon 1 31425
2 salmon 2 324116
3 salmon 3 676852Using filter() to exclude rows also drops NAs!
The problem with using filter() to exclude
To properly use filter(), we would need to do something like:
# A tibble: 4 × 3
species region losses
<chr> <chr> <dbl>
1 salmon 1 31425
2 salmon 2 324116
3 salmon 3 676852
4 salmon 3 NA
Work smarter with dplyr 1.2.0!
New filter_out() function
Use…
⬢ filter() to keep rows
⬢ filter_out() to drop rows
Work smarter with filter_out()
Now, we just have to run:
# A tibble: 4 × 3
species region losses
<chr> <chr> <dbl>
1 salmon 1 31425
2 salmon 2 324116
3 salmon 3 676852
4 salmon 3 NA
when_any() and when_all()
Issues with using filter() and |
Let’s look at this sample dataset:
Issues with using filter() and |
Keep rows where region 7 or 8 have losses over 400,000 OR and where regions 2 or 9 have losses over 300,000.
# A tibble: 8 × 3
# Groups: region [4]
species region losses
<chr> <chr> <dbl>
1 salmon 2 324116
2 salmon 2 197239
3 salmon 7 231487
4 salmon 7 475115
5 salmon 8 442659
6 salmon 8 327323
7 salmon 9 311127
8 salmon 9 286601



Issues with using filter() and |
Keep rows where region 7 or 8 have losses over 400,000 OR and where regions 2 or 9 have losses over 300,000. In this case, Rows 1, 4, 5, and 7)
# A tibble: 4 × 3
# Groups: region [4]
species region losses
<chr> <chr> <dbl>
1 salmon 2 324116
2 salmon 7 475115
3 salmon 8 442659
4 salmon 9 311127
Work smarter with dplyr 1.2.0!
New when_any() and when_all() functions
Use…
⬢ when_any() to specify “or” conditions
⬢ when_all() to specify “all” conditions
Work smarter with filter() + when_any()
# A tibble: 4 × 3
# Groups: region [4]
species region losses
<chr> <chr> <dbl>
1 salmon 2 324116
2 salmon 7 475115
3 salmon 8 442659
4 salmon 9 311127
Work smarter with filter() + when_all()
# A tibble: 2 × 3
# Groups: region [2]
species region losses
<chr> <chr> <dbl>
1 salmon 7 475115
2 salmon 8 442659
New recoding functions
Recoding has always been a pain
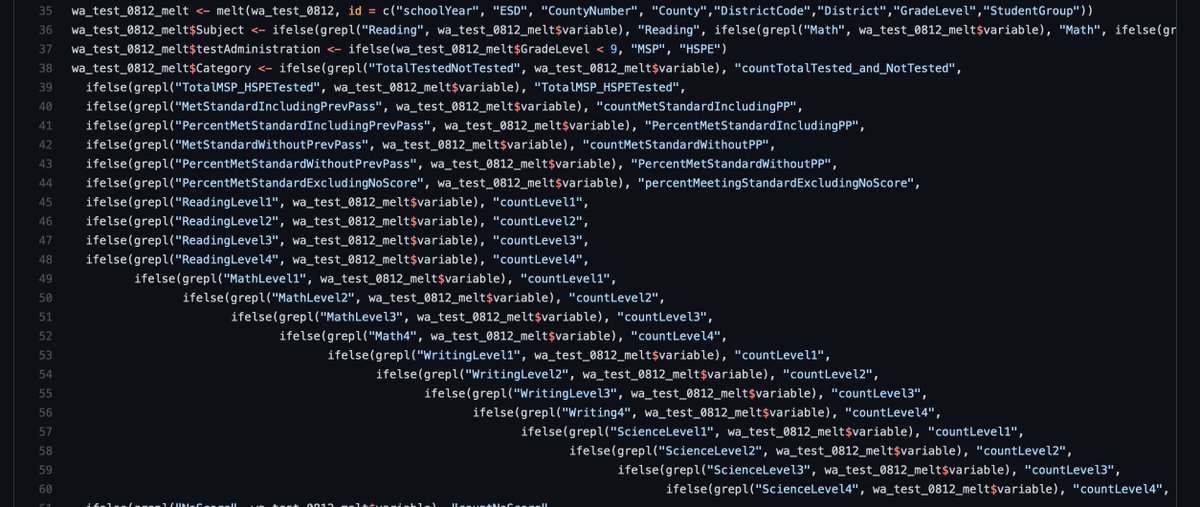
Recoding with case_when()
monthly_losses_data |>
select(species, geo_group, region, losses) |>
filter(geo_group == "area") |>
mutate(
production_area =
case_when(
region == "1" ~ "Jæren",
region == "2" ~ "Ryfylke",
region == "3" ~ "Sotra",
region == "4" ~ "Stadt",
region == "5" ~ "Hustadvika",
region == "6" ~ "Nordmøre",
region == "7" ~ "Nord-Trøndelag",
region == "8" ~ "Bodø",
region == "9" ~ "Vestlfjorden",
region == "10" ~ "Andfjorden",
region == "11" ~ "Kvaløya",
region == "12" ~ "Vest-Finnmark",
region == "13" ~ "Øst-Finnmark",
.default = NA_character_
)
)Recoding with case_when()
# A tibble: 1,512 × 5
species geo_group region losses production_area
<chr> <chr> <chr> <dbl> <chr>
1 salmon area 1 31425 Jæren
2 salmon area 2 324116 Ryfylke
3 salmon area 3 844829 Sotra
4 salmon area 4 676852 Stadt
5 salmon area 5 109269 Hustadvika
6 salmon area 6 548921 Nordmøre
7 salmon area 7 231487 Nord-Trøndelag
8 salmon area 8 442659 Bodø
9 salmon area 9 311127 Vestlfjorden
10 salmon area 10 288849 Andfjorden
# ℹ 1,502 more rowsRecoding with recode()
monthly_losses_data |>
select(species, geo_group, region, losses) |>
filter(geo_group == "area") |>
mutate(
production_area =
recode(
region,
"1" = "Jæren",
"2" = "Ryfylke",
"3" = "Sotra",
"4" = "Stadt",
"5" = "Hustadvika",
"6" = "Nordmøre",
"7" = "Nord-Trøndelag",
"8" = "Bodø",
"9" = "Vestlfjorden",
"10" = "Andfjorden",
"11" = "Kvaløya",
"12" = "Vest-Finnmark",
"13" = "Øst-Finnmark",
.default = NA_character_
)
)Recoding with recode()
# A tibble: 1,512 × 5
species geo_group region losses production_area
<chr> <chr> <chr> <dbl> <chr>
1 salmon area 1 31425 Jæren
2 salmon area 2 324116 Ryfylke
3 salmon area 3 844829 Sotra
4 salmon area 4 676852 Stadt
5 salmon area 5 109269 Hustadvika
6 salmon area 6 548921 Nordmøre
7 salmon area 7 231487 Nord-Trøndelag
8 salmon area 8 442659 Bodø
9 salmon area 9 311127 Vestlfjorden
10 salmon area 10 288849 Andfjorden
# ℹ 1,502 more rowsRecoding with recode() + rlang
Say we map our areas to production areas (POs):
Recoding with recode() + rlang
We can use rlang’s !!! to splice the list and use it in recode():
Recoding with recode() + rlang
# A tibble: 1,512 × 5
species geo_group region losses production_area
<chr> <chr> <chr> <dbl> <chr>
1 salmon area 1 31425 Jæren
2 salmon area 2 324116 Ryfylke
3 salmon area 3 844829 Sotra
4 salmon area 4 676852 Stadt
5 salmon area 5 109269 Hustadvika
6 salmon area 6 548921 Nordmøre
7 salmon area 7 231487 Nord-Trøndelag
8 salmon area 8 442659 Bodø
9 salmon area 9 311127 Vestlfjorden
10 salmon area 10 288849 Andfjorden
# ℹ 1,502 more rowsWork smarter with dplyr 1.2.0!
Recoding vs replacing
⬢ recoding is creating an entirely new column using values from an existing column
⬢ replacing is partially updating an existing column with new values
New recoding and replacing functions
Recall case_when():
# A tibble: 4 × 4
species region losses loss_rating
<chr> <chr> <dbl> <chr>
1 salmon 1 31425 Low
2 salmon 2 324116 High
3 salmon 3 844829 High
4 salmon 4 676852 High New recoding and replacing functions
Three new functions to join the case_when() family. Use:
⬢ case_when() when recoding and matching with conditions
⬢ recode_values() when recoding and matching with values
⬢ replace_when() when replacing and matching with conditions
⬢ replace_values() when replacing and matching with values
New recode_values() function
Recall that our region is a value and we would like to recode our dataset
monthly_losses_data |>
select(species, geo_group, region, losses) |>
filter(geo_group == "area") |>
mutate(
production_area = region |>
recode_values(
"1" ~ "Jæren",
"2" ~ "Ryfylke",
"3" ~ "Sotra",
"4" ~ "Stadt",
"5" ~ "Hustadvika",
"6" ~ "Nordmøre",
"7" ~ "Nord-Trøndelag",
"8" ~ "Bodø",
"9" ~ "Vestlfjorden",
"10" ~ "Andfjorden",
"11" ~ "Kvaløya",
"12" ~ "Vest-Finnmark",
"13" ~ "Øst-Finnmark"
)
)New recode_values() function
Recall that our region is a value and we would like to recode our dataset
monthly_losses_data |>
select(species, geo_group, region, losses) |>
filter(geo_group == "area") |>
mutate(
production_area = region |>
recode_values(
"1" ~ "Jæren",
"2" ~ "Ryfylke",
"3" ~ "Sotra",
"4" ~ "Stadt",
"5" ~ "Hustadvika",
"6" ~ "Nordmøre",
"7" ~ "Nord-Trøndelag",
"8" ~ "Bodø",
"9" ~ "Vestlfjorden",
"10" ~ "Andfjorden",
"11" ~ "Kvaløya",
"12" ~ "Vest-Finnmark",
"13" ~ "Øst-Finnmark",
unmatched = "error"
)
)New recode_values() function
# A tibble: 1,512 × 5
species geo_group region losses production_area
<chr> <chr> <chr> <dbl> <chr>
1 salmon area 1 31425 Jæren
2 salmon area 2 324116 Ryfylke
3 salmon area 3 844829 Sotra
4 salmon area 4 676852 Stadt
5 salmon area 5 109269 Hustadvika
6 salmon area 6 548921 Nordmøre
7 salmon area 7 231487 Nord-Trøndelag
8 salmon area 8 442659 Bodø
9 salmon area 9 311127 Vestlfjorden
10 salmon area 10 288849 Andfjorden
# ℹ 1,502 more rowsUsing recode_values() and a lookup table
Let’s create a lookup table from our list:
# A tibble: 13 × 2
from to
<chr> <chr>
1 1 Jæren
2 2 Ryfylke
3 3 Sotra
4 4 Stadt
5 5 Hustadvika
6 6 Nordmøre
7 7 Nord-Trøndelag
8 8 Bodø
9 9 Vestlfjorden
10 10 Andfjorden
11 11 Kvaløya
12 12 Vest-Finnmark
13 13 Øst-Finnmark Using recode_values() and a lookup table
We can use a lookup to recode our values:
Using recode_values() and a lookup table
# A tibble: 1,512 × 5
species geo_group region losses production_area
<chr> <chr> <chr> <dbl> <chr>
1 salmon area 1 31425 Jæren
2 salmon area 2 324116 Ryfylke
3 salmon area 3 844829 Sotra
4 salmon area 4 676852 Stadt
5 salmon area 5 109269 Hustadvika
6 salmon area 6 548921 Nordmøre
7 salmon area 7 231487 Nord-Trøndelag
8 salmon area 8 442659 Bodø
9 salmon area 9 311127 Vestlfjorden
10 salmon area 10 288849 Andfjorden
# ℹ 1,502 more rowsNew replace_values() function
Using replace_values() to fix some names:
New replace_values() function
# A tibble: 1,512 × 5
species geo_group region losses production_area
<chr> <chr> <chr> <dbl> <chr>
1 salmon area 1 31425 Jæren
2 salmon area 2 324116 Ryfylke
3 salmon area 3 844829 Sotra
4 salmon area 4 676852 Stadt
5 salmon area 5 109269 Hustadvika
6 salmon area 6 548921 Nordmøre + Sør-Trøndelag
7 salmon area 7 231487 Nord-Trøndelag + Bindal
8 salmon area 8 442659 Bodø
9 salmon area 9 311127 Vestlfjorden
10 salmon area 10 288849 Andfjorden
# ℹ 1,502 more rowsNew replace_when() function
case_when() requires .default, otherwise it will default to NA:
# A tibble: 4 × 4
species region losses loss_rating
<chr> <chr> <dbl> <dbl>
1 salmon 1 31425 31425
2 salmon 2 324116 324116
3 salmon 3 844829 500000
4 salmon 4 676852 500000New replace_when() function
replace_when() knows you will only replace some of the values, so a .default is not necessary:
Additional thoughts
case_match() has been soft deprecated

Updating your LLM
- Paste documentation into the conversation
- Add a CLAUDE.md file to your project
- Point it to the docs
- Use an MCP server
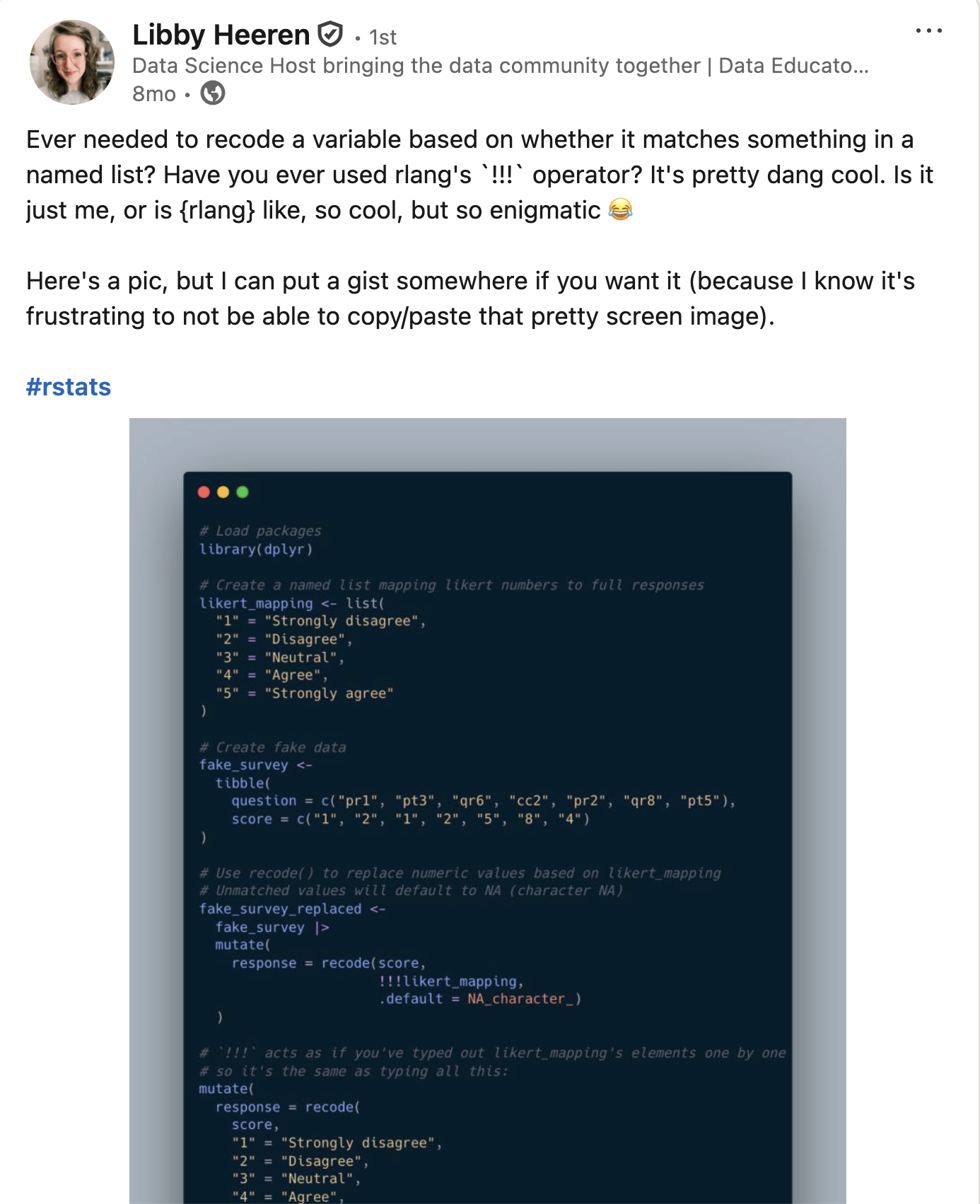
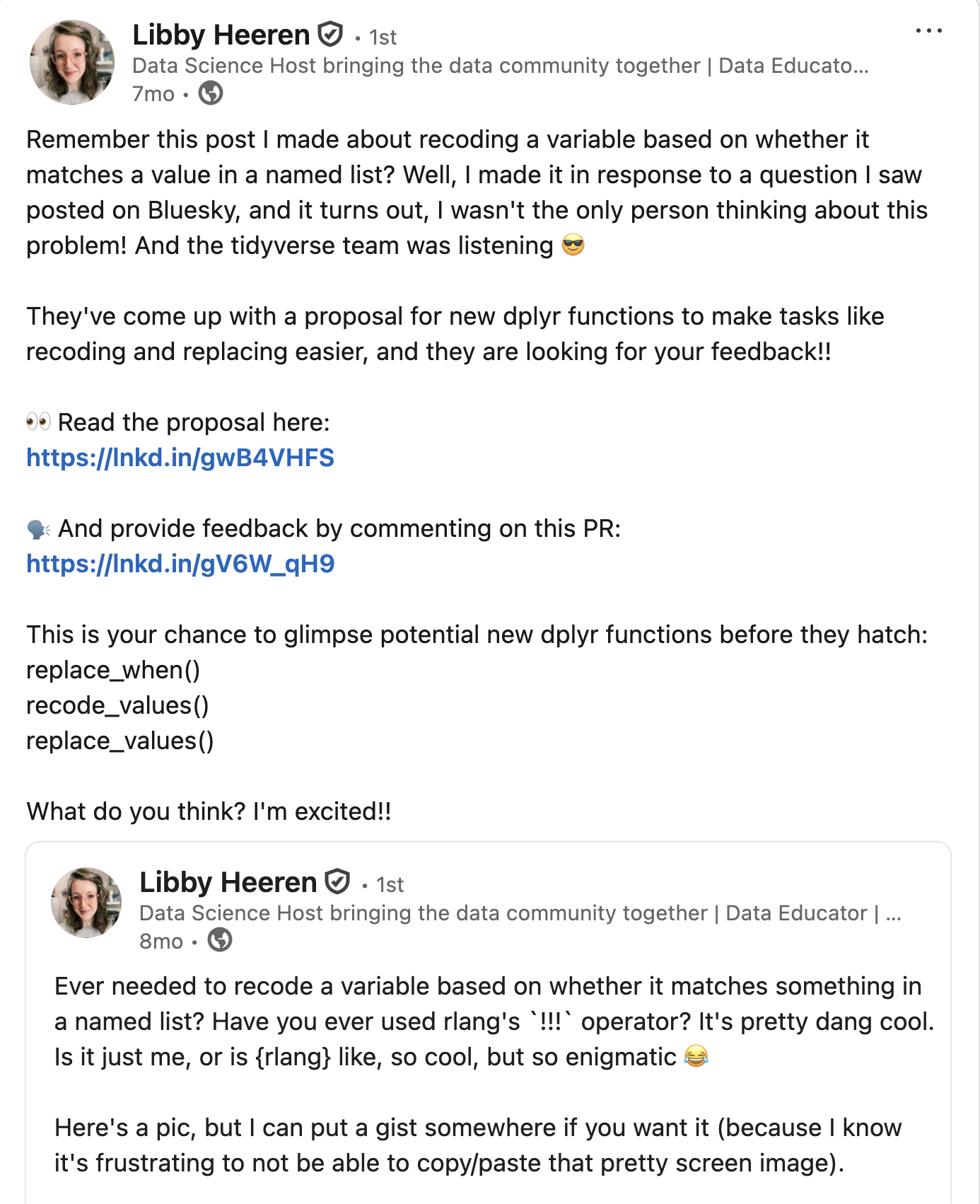
Community feedback is sooo important
We are looking for #rstats community feedback on 3 new dplyr functions!
We're aiming to expand the
filter()family:Read more and leave feedback here: github.com/tidyverse/ti…
filter()to keep rowsfilter_out()to drop rowswhen_any()andwhen_all()as modifiers
[image or embed] — Davis Vaughan (@davisvaughan.bsky.social) November 7, 2025 at 10:03 AM
Community feedback is sooo important
Summary
New dplyr 1.2.0 functions
⬢ filter_out(), the missing complement to filter(), and accompanying when_any() and when_all() helpers
⬢ recode_values(), replace_values(), and replace_when(), three new functions for recoding and replacing
Look out for the upcoming Data Science Lab recording!
Installing dplyr 1.2.0
Upgrade today!

Thank you
Acknowledgements
- Davis Vaughan and the tidyverse team
- Libby Heeren for her awesome example
- Emojis from OpenMoji
- Photo from Unsplash
Links
ivelasq-dplyr-1-2-0.share.connect.posit.cloud